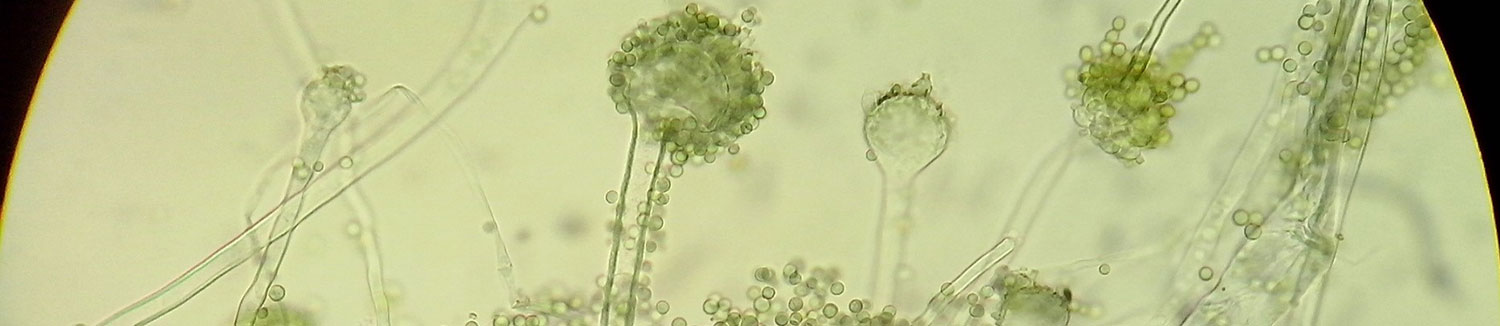
Fungal Armory and Weaponry: A Source for Pharmaceutical Development
The Land O’ Lakes conference series strives to be at the forefront of pharmaceutical innovation. Our Monday evening reception dinners feature speakers with a unique perspective. For our September (DMPK) Land O’ Lakes Conference we had UW Madison‘s own Nancy Keller, PhD share the latest updates and insights from her research.
Professor Keller works in the Department of Medical Microbiology and Immunology. The Keller Lab explores the genetic regulation of virulence and natural product synthesis by fungi. Currently, her research focuses on Aspergillus spp. Working alongside her students; she tries to dissect the genetic aspects of Aspergillus spp that can be harnessed to develop useful biological products.
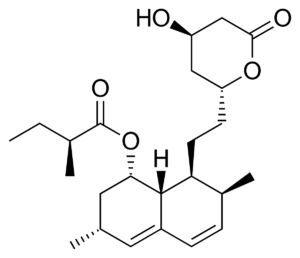
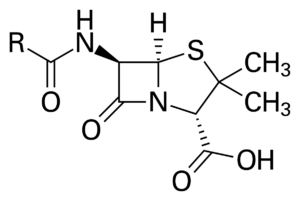
Microorganisms like fungi are natural product synthetic chemists. Professor Keller began her talk by explaining how modern medicine is already using fungal metabolites. The well-known antibiotic Penicillin was first discovered and derived from the Penicillium fungi. The cholesterol-reducing drug Lovastatin was derived from Aspergillus terreus. Foods like mushrooms and fermented products have statin compounds with cholesterol-reducing abilities.
 |
 |
Fungal Metabolites: Sources of Fungal Armory and Weaponry
Penicillin and Lovastatin are thought to be just the first few useful metabolites. There are many metabolites yet to be discovered and extracted from fungi. Professor Keller described three categories of fungal metabolites based on where, why and how they are made in the fungus.
- Tissue Specificity/Development
- Offense/Defense
- Self-Protection
Tissue Specificity/Development
 Fungi produce natural products in specific tissues for ecological fitness. For example, many fungi produce various melanins in their spores to protect the spores from UV radiation. Some studies are exploring the possibility of using fungal melanins in sunscreens. For example, Aspergillus nidulans expresses a melanin precursor called DHICA (5, 6- Dihydroxyindole-2-carboxylic acid). DHICA was tested on the skin of mice and it was equivalent to t a sun protective factor (SPF) 20 sunscreen. The study concluded that DHICA has the potential to be a UVB protectant 1.
Fungi produce natural products in specific tissues for ecological fitness. For example, many fungi produce various melanins in their spores to protect the spores from UV radiation. Some studies are exploring the possibility of using fungal melanins in sunscreens. For example, Aspergillus nidulans expresses a melanin precursor called DHICA (5, 6- Dihydroxyindole-2-carboxylic acid). DHICA was tested on the skin of mice and it was equivalent to t a sun protective factor (SPF) 20 sunscreen. The study concluded that DHICA has the potential to be a UVB protectant 1.
Another potential protectant from UV rays is (3R)-5-hydroxymellein, a secondary metabolite produced by lichens. Testing on mouse cell lines showed that (3R)-5-hydroxymellein recovered damage caused by UVB radiation and inhibited melanin synthesis. This study concluded that (3R)-5-hydroxymellein could be used to develop multifunctional skin protectants 2.
Emodin is a plant-derived metabolite. While found in rhubarb and buckthorn, it is also found in several fungi like Aspergillus sp, Pyrenochaeta, and Pestaloitiopsis. Emodin is commonly used in laxatives but has other beneficial properties as well. It has an anti-cancer effect on many cancers including pancreatic cancer. It can block cytomegalovirus and herpes simplex infections. It may also be able to address obesity in by inhibiting enzyme 11beta-HSD1 < 3.
Offense/Defense
Many fungi and bacteria are at odds with each other in nature, competing for common resources, and adapting to prevail. This is why some bacteria can produce good antifungals. It also means that fungi are a potent source of antibacterials. Penicillin is the most well-known fungal antibacterial.
Bacterial and fungal infections are a risk in farming as well. Ralstonia solanacearum is an aerobic bacterium that is found in soils. Ralstonia can infect and ruin many crops including potato, tomato, eggplant, banana, ginger, tobacco, pepper, olive, rose, and soybeans by causing wilt disease. The fungus Fusarium spp also causes wilt disease. and tends to be in the same soils and impacts the same crops as Ralstonia.
Ralstonia secretes a compound that causes Fusarium spp to develop chlamydospores. The bacteria invade these hardy spores to survive winter weather. To combat the bacteria, Fusarium spp produces the metabolite Bikaverin. Bikaverin inhibits and represses Ralstonia growth 4.
The understanding of how soil microbes interact is still in its infancy. Professor Keller believes that understanding this interaction can help us to identify antibacterials and antifungals that may someday not only help protect crops from disease but also result in the finding of useful pharmaceuticals.
Self-Protection
Many of the drugs that fungi make are also toxic to other fungi. Therefore some fungi contain a self-protection gene that shields them from their own toxin. Two examples are the proteasome inhibitor fellutamide and the methionine aminopeptidase inhibitor fumagillin.
Proteasome inhibitors are among the newest drugs being studied in anti-cancer therapy. They are being used to treat multiple myeloma and lymphoma. Scientists believe that proteasome inhibitors are one key drug to combating these types of cancer. Fellutamide is a powerful proteasome inhibitor derived from several fungi including Aspergillus nidulans. Because fungi contain the same proteasome proteins as humans, fellutamide is also toxic to fungi. Professor Keller and colleagues found that the fungus A. nidulans contains a self-protection mechanism that protects it from its own fellutamide 5.
Fumagillin is another biomolecule and antimicrobial agent. It was originally isolated from Aspergillus fumigatus in 1949 and in 2013 Professor Keller’s lab identified the fumagillin gene cluster in A. fumigatus and found that the fungus contained self-protection genes in the gene cluster 6. Fumagillin has been used to treat Microsporidiosis for several decades. It is also used to treat microsporidian parasites in honeybees and fish. However, Fumagillin may also be used for the treatment of cancer and rheumatoid arthritis. Fumagillin targets methionine aminopeptidase 2 (METAP2) an enzyme that promotes angiogenesis. Angiogenesis is the generation and growth of new blood vessels. Solid state tumors and rheumatoid arthritis spread via angiogenesis. By inhibiting angiogenesis, Fumagillin may help prevent the spread of cancer tumors. The Fumagillin treatment for cancers is currently in clinical trials and studies are underway for its efficacy in treating rheumatoid arthritis.
Unlocking the treasure chest
Clearly, fungal metabolites and biomolecules have many useful applications. However, there are many challenges in harnessing these natural compounds for our benefit. It is not always evident which fungal compound is causing the desired results. The compound needs to be identified and the corresponding fungal genes sequenced. Once this is done the final challenge is to produce the desired natural products on a much larger scale.
 There are three different approaches to do this.
There are three different approaches to do this.
Native Production
The first approach is to coax the native strains to ramp up production. This includes methods such as genome mining, genetic manipulation, non-standard culturing and mutagenesis.
Synthetic Biology
The second method is synthetic biology. This involves creating biosynthetic pathways and cell-free expression systems to synthesize the desired compound.
Hetereologous Expression
The third and final approach is heterologous expression. This is the method used and studied extensively at the Keller Lab. Heterologous expression involves the expression of a gene or gene fragment in a host organism that does not normally produce the gene or gene fragment. Professor Keller explained how they have used heterologous expression in her lab.
The Keller lab and colleagues use Fungal Artificial Chromosomes (FACs) to mine fungal secondary metabolomes. Fungal genomes are unbiasedly sheared to create a FAC library. The FACs are then inserted and edited in E.coli where genetic deletions can take place. Once the FAC creation and recombination is done, Aspergillus nidulans is used as the host for heterologous expression. Finally, High-Resolution Liquid Chromatography-Mass Spectrometry (LC-MS) is used to identify secondary metabolomes.
Each LC-FTMS analysis resulted in the detection of approximately 10,000 compounds. The “FAC-Score” uses a decoy approach to identify likely FAC products. Using this approach they were able to detect gene cluster products from 15/56 FAC-transformed strains.
Professor Keller and her colleagues published their study “A Scalable Platform to Identify Fungal Secondary Metabolites and Their Gene Clusters” in the August 2017 Volume 13 No 8 edition of the magazine nature chemical biology. You can review the complete article here.
Professor Keller’s presentation gave a brief look at a new and growing opportunity. Fungi represent an exciting frontier for fighting human diseases. Professor Keller and her lab are using multiple technologies to leverage their discoveries to enable us to harness fungi as warriors battling for successful human therapies of the future.
Check out our Biologics curriculum to expand your knowledge on the development of Biological pharmaceuticals. Attend our Land O’ Lakes Conferences to connect with researchers like Nancy Keller and others in the pharmaceutical industry.
Sources:
Marine fungal DHICA as a UVB protectant: Assessment under in vitro and in vivo conditions; Shanuja SK, Iswarya S, Gnanamani A. (https://www.ncbi.nlm.nih.gov/pubmed/29367149)
A Multifunctional and Possible Skin UV Protectant, (3R)-5-Hydroxymellein, Produced by an Endolichenic Fungus Isolated from Parmotrema austrosinense; Zhao L, Kim JC, Paik MJ, Lee W, Hur JS (https://www.ncbi.nlm.nih.gov/pubmed/28035958)
Anti-Cytomegalovirus Activity of the Anthraquinone Atanyl Blue PRL; Zohaib Alam, Zainab Al-Mahdi, Yali Zhu, Zachary McKee, Deborah S Parris, Hardik I. Parikh, Glen E. Kellogg, Alison Kuchta, Michael A McVoy (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4289655/)
Conserved Responses in a War of Small Molecules between a Plant-Pathogenic Bacterium and Fungi; Joseph E. Spraker, Philipp Wiemann, Joshua A. Baccile, Nandhitha Venkatesh, Julia Schumacher, Frank C. Schroeder, Laura M. Sanchez, Nancy P. Keller (https://mbio.asm.org/content/9/3/e00820-18)
Resistance Gene-Guided Genome Mining: Serial Promoter Exchanges in Aspergillus nidulans Reveal the Biosynthetic Pathway for Fellutamide B, a Proteasome Inhibitor; Yeh HH, Ahuja M, Chiang YM, Oakley CE, Moore S, Yoon O, Hajovsky H, Bok JW, Keller NP, Wang CC, Oakley BR (https://www.ncbi.nlm.nih.gov/pubmed/27294372)
Prototype of an intertwined secondary-metabolite supercluster; Wiemann P, Guo CJ, Palmer JM, Sekonyela R, Wang CC, Keller NP (https://www.ncbi.nlm.nih.gov/pubmed/24082142)